Welcome to the Barrett lab in the Department of Biology at McGill University! Our research uses approaches in molecular ecology, population genomics and experimental evolution to ask questions about the interactions between ecological and evolutionary processes.
Our purpose is to discover new and interesting things in biology, and to create a collaborative, productive, and friendly environment in which we can train future generations of scientists.
Lab Members

Rowan Barrett
Principal Investigator
Rowan is the Canada Research Chair of Biodiversity Science at McGill University. He completed his M.Sc. at McGill in 2005, conducting experimental evolution with microbes. He completed his Ph.D. at the University of British Columbia in 2010, where he studied the genetics of adaptation in threespine stickleback. He then moved to Harvard University as a Howard Alper postdoctoral fellow, where he investigated ecological genomics using deer mice. Since 2013 Rowan has been a professor in the Redpath Museum and Department of Biology at McGill. He is broadly interested in the reciprocal interactions between ecological and evolutionary processes, and the mechanisms by which these forces impact genomic variation in natural populations.
Please click on each member to see their personal website

Mary Kathleen (MK) Hickox
Graduate Student

Ananda Martins
Graduate Student

Eric Wootton
Graduate Student

Laura Lardinois
Graduate Student

Akhil Kholwadwala
Graduate Student

Lucas Eckert
Graduate Student

Paige Smallman
Graduate Student

Tim Gemeinhardt
Graduate Student

Tim Thurman
Graduate Student

Sara Wuitchik
Graduate Student

Juntao Hu
Graduate Student

Dieta Hanson
Graduate Student

Arash Askary
Graduate Student

Sara Kurland
Graduate Student

Bushra Sial
Graduate Student

Madlen Stange
Postdoctoral Fellow

Antoine Paccard
Postdoctoral Fellow

Caroline LeBlond
Lab Manager

Juliette Lemoine (BIOL 479)
Undergraduate Student

Avery Albert
Undergraduate Student

Kiran Yendamuri
Undergraduate Student

Samantha LaPenna
Undergraduate Student

Emily Brown
Undergraduate Student

Nicole Stinson
Undergraduate Student

Rachel Takasaki
Undergraduate Student

Michael Maddalena
Undergraduate Student

Lauren Bennett
Undergraduate Student

Scarlett (Yiyi) Xiao
Undergraduate Student

Frida-Cecilia Acosta-Parenteau
Undergraduate Student (BIOL 466)

Savannah Bissegger-O’Connor
Undergraduate Student (BIOL 466)

Olivia Fraser Barsby
Undergraduate Student

Natalie Warren
Undergraduate Student

Jaren Mari Abergas
Undergraduate Student

Mathilde Salamon
Graduate Student

Alan Garcia-Elfring
Graduate Student

Maxime Guglielmetti
Undergraduate Student

Linda Tang
Undergraduate Student

Brook Brown
Undergraduate Student
Johanna Arnet
Undergraduate Student

Åsa Lind
Research Associate

Marc-Olivier Beausoleil
Graduate Student

Alina Shimizu-Jozi
Undergraduate Student

Charles Xu
Graduate Student

Andrew Kemp
Undergraduate Student

Eva Savard
Undergraduate Student

Victoria Glynn
Graduate Student

Janay Fox
Graduate Student

Marcus Lin
Graduate Student
Research
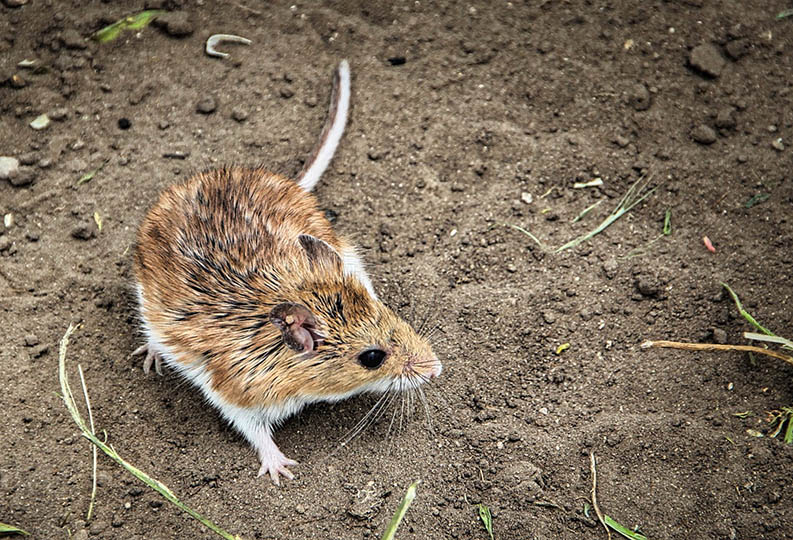
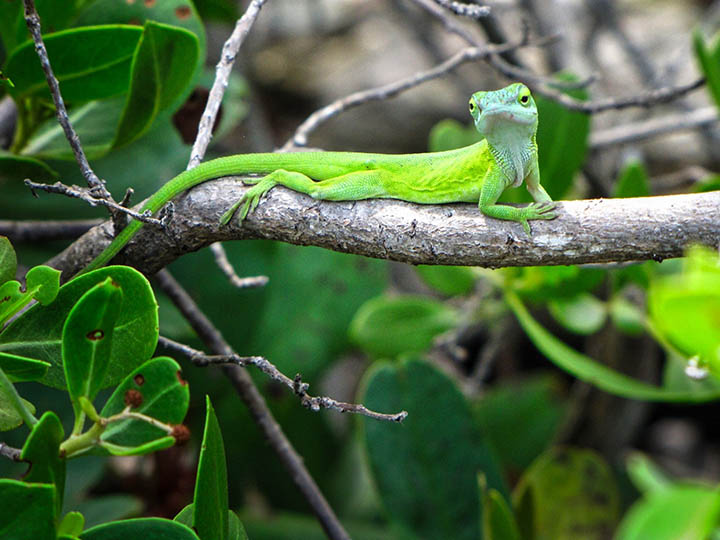
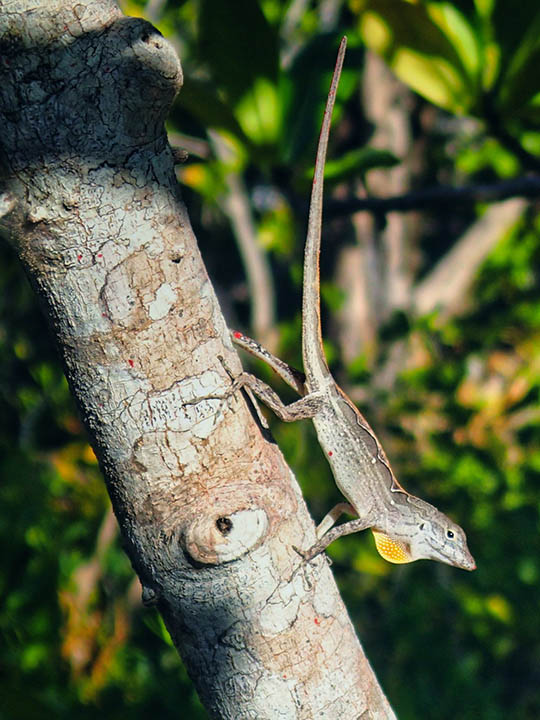
Rapid environmental change results in elevated extinction rates and thus poses a major threat to biodiversity. One way in which organisms might respond to these changes – and thus persist into the future – is through adaptive evolution. Our research investigates this possibility by connecting traits related to Darwinian fitness (i.e., survival and reproductive success) with their underlying genetic architecture in relevant ecological scenarios. The extent to which evolution is predictable is a major unresolved question in biology, and crucial for understanding how populations will evolve in response to rapidly changing environmental conditions. Although natural selection is a deterministic process, the predictability of evolution might be limited because the ecological sources of selection and the genetic basis of adaptation can be complex. Our research combines a variety of approaches and study systems to help understand this complexity. We generate and test hypotheses about the predictability of evolution through a combination of ecological field experiments, molecular biology, genomics, and computational biology. Our main study systems are threespine stickleback fish (Gasterosteus aculeatus), deer mice (Peromyscus maniculatus), and anolis lizards (A. sagrei and A. carolinensis), but we often work with other organisms too (such as bacteria, Galapagos finches, or Heliconius butterflies). We aim to quantify the contributions of genome-wide genetic variation to fitness, and to understand the ecological and evolutionary forces that have shaped these patterns of variation between individuals, populations, and closely related species.
You can find code that we have developed for our research on our lab GitHub page here.
Publications
Lab member names are underlined
"*" indicates equal contributions
2025
He, X., Maruki, T., Morgado-Gamero, W.B., Barrett, R.D.H., Fugère, V., Fussmann G.F., Gonzalez, A., Shapiro, B.J., Cristescu, M.E. (2025). Environmental RNA-Based Metatranscriptomics as a Novel Biomonitoring Tool: A Case Study of Glyphosate-Based Herbicide Effects on Freshwater Eukaryotic Communities. Molecular Ecology. https://doi.org/10.1111/mec.70164. |
|
| Ho, W.-Z., Lind, Å.J., Smith, R., Dwyer-Samuel, F., Pilgrim, S., Gear, G., Laing, R., Tomy, G., Mallory, M.L., Enook, J., Zahaby, Y., Provencher J.F., Barrett, R.D.H. (2025) Epigenetic responses to anthropogenic versus natural sources of oil exposure differ in wild Arctic seabird populations. Evolutionary Applications doi.org/10.1111/eva70125. | |
| Fox, J., Reader, S., Guigueno, M.F., Barrett, R.D.H. (2025) Developmental Behavioural Plasticity and DNA Methylation Patterns in Response to Predation Stress in Trinidadian Guppies. Molecular Ecology 0.e:17831. doi.org/10.1111/mec.17831. | |
| Glynn, V.M., Marangoni, L.F., Guglielmetti, M., Tapia, E.R., Ali, V., Quintero, H., Guerra, E.C.R., Yuval, M., Kline, D.I., Leray, M., Connolly, S.R., Barrett, R.D.H. (2025) The role of holobiont composition and environmental history in thermotolerance of Tropical Eastern Pacific corals. Current Biology 35: 1-16.e1-e7. | |
| Fox, J., Reader, S., Barrett, R.D.H. (2025) Rapid neural DNA methylation responses to predation stress in Trinidadian guppies. Molecular Ecology 0:e17774. doi.org/10.1111/mec.17774. | |
| Garcia-Elfring, A., Roffey, H.L., Abergas, J.M., Hendry, A.P., Barrett, R.D.H. (2025) GTP cyclohydolase 1 and axanthism in ball pythons: a new vertebrate model for pterin-based pigmentation. Animal Genetics 56: e70011. https://doi.org/10.1111/age.70011. | |
| Carrión, P.L., Beausoleil, M., Raeymaekers, J.A.M., De León, L.F, Chaves, J.A., Sharpe, D.M.T., Huber, S.K., Herrel, A., Gotanda, K.M., Koop, J.A.H., Knutie, S.A., Clayton, D.H., Podos, J., Barrett, R.D.H., Guichard, F., Hendry, A.P. (2025) Darwin’s finches and climate change: insights from a resilient system. Journal of Evolutionary Biology doi.org/10.1093/jeb/voaf138 | |
| Katkov, E. Gros, M., Fugère, V., Sychterz, A., Tadiri, C.P., Barrett, R.D.H., Cristescu, M., Fussmann, G.F., Gonzalez, A. (2025) Connectivity and nutrient enrichment affect the productivity and stability of aquatic meta-ecosystems. Proceedings of the Royal Society of London B doi.org/10.1098/rspb.2025.1197 | |
| Eckert, L., Eckert, I., Rahn, O., So, C.P., Barrett, R.D.H. (2025) Using herbarium collections to study genetic responses to global change. New Phytologist doi.org/10.1111/nph.70454. | |
| Zhao, J., Luo, M., Merila, J., Barrett, R.D.H., Guo, B., Hu, J. (2025) The interplay between epigenomic and transcriptomic variation in divergence of stickleback ecotypes. BMC Biology 23(1):70. doi.org/10.1186/s12915-025-02176-0. | |
| Xu, C. C. Y., Fugère, V., Barbosa da Costa, N., Beisner, B. E., Bell, G., Cristescu, M. E., Fussmann, G. F., Gonzalez, A., Shapiro, B. J., Barrett, R. D. H. (2025). Pre-exposure to stress reduces loss of community and genetic diversity following severe environmental disturbance. Current Biology. https://doi.org/10.1016/j.cub.2025.01.037 | |
| Loria, A., Tournayre, O., Hébert, M.-P., da Costa, N.B., Fugère, V., Barrett, R.D.H., Beisner, B.E., Gonzalez, A. and Cristescu, M.E. (2025), Estimating Rapid Diversity Changes During Acute Herbicide Contamination Using Environmental DNA. Environmental DNA, 7: e70029. https://doi.org/10.1002/edn3.70029 | |
| Hu, J., Ren, X., Barrett, R.D.H., Samma, E., Sulkowski, M., Brady, S.P. (2025) Alternative splicing and parallel patterns of gene expression underlie population and life history divergence in an amphibian. Ecology and Evolution 15(11): e72481. doi.org/10.1002/ece3.72481 | |
| Garcia-Elfring, A., Roffey, H.L., Abergas, J.M., Hendry, A.P., Tzika, A.C., Barrett, R.D.H. (2025) Ball python colour morphs implicate mc1r in skin color patterning. Pigment Cell and Melanoma Research doi.org/10.1111.pcmr13215. |
2024
| Salamon, M., Astorg, L., Paccard, A., Chain, F., Hendry, A.P., Derry, A.M.*, Barrett, R.D.H.* (2024) Limited migration from physiological refugia constrains the rescue of native gastropods facing an invasive predator. Evolutionary Applications 17(10): e70004. https://doi.org/10.1111/eva.70004 | |
| Derrick, E., Costa, N.B., Barrett, R.D.H., Shapiro, B.J. A glyphosate-based herbicide selects for genetic changes while retaining within-species diversity in a freshwater bacterioplankton community. The ISME Journal submitted. https://www.biorxiv.org/content/10.1101/2024.09.17.613573v1.full.pdf. | |
| Baduel, P., Sammarco, I., Barrett, R.D.H., Coronado-Zamora, M., Crespel, A., Díez-Rodríguez, B., Fox, J., Galanti, D., González, J., Jueterbock, A., Wootton, E., & Harney, E. (2024) The evolutionary consequences of interactions between the epigenome, the genome and the environment. Evolutionary Applications http://dx.doi.org/10.1111/eva.13730 | |
| Fox, J., Toure, M.W., Heckley, A., Raina, F., Reader, S., Barrett, R.D.H. (2024) Insights into adaptive behavioural plasticity from the guppy model system. Proceedings of the Royal Society of London B https://doi.org/10.1098/rspb.2023.2625 | |
| Fox, J., Hunt, D., Hendry, A.P., Chapman, L., Barrett, R.D.H. (2024) Counter-gradient variation contributes to divergence of colonizing populations of fish in varying dissolved oxygen environments. Molecular Ecology http://doi.org/10.1111/mec.17419. | |
| Hendry, A.P., Barrett, R.D.H., Bell, A., Bell, M., Bolnick, D.I., Gotanda, K., Haines, G., Lind, Å., Packer, M., Peichel, C.L., Peterson, C., Poore, H., Massengill, R., McClellan, K., Steinel, N.C., Sanderson, S., Walsh, M., Weber, J.N., Derry, A.M. (2024) Designing eco-evolutionary experiments restoration projects: opportunities and constraints revealed during stickleback introductions. Ecology and Evolution https://doi.org/10.1002/ece3.11503. |
2023
| Beausoleil, M., Carrion-Aviles, P.L., Podos, J., Camacho, C., Richard, R., Lalla, K., Raeymaekers, J.A.M., Knutie, S.A., De Leon, L.F., Huber, S.K., Chaves, J.A., Clayton, D.H., Koop, J.A.H., Sharpe, D., Gotanda, K.M., Barrett, R.D.H., Hendry, A.P. (2023) The fitness landscape of a community of Darwin’s finches. Evolution https://doi.org/10.1093/evolut/qpad160. | |
| Martins, A.R.P., Warren, N., McMillan, W.O., Barrett, R.D.H. (2023) Spatiotemporal dynamics in butterfly hybrid zones. Insect Science 10.1111/1744-7917.13262. | |
| Xu, C.C.Y., Lemoine, J., Albert, A., Mac Whirter, É, Barrett, R.D.H. (2023) Community assembly of the human piercing microbiome. Proceedings of the Royal Society of London B. https://doi.org/10.1098/rspb.2023.1174. | |
| Monchamp, M.-E., Taranu, Z.E., Garner, R.E., Rehill, T., Morissette, O., Iversen, L.L., Fugère, V., Littlefair, J.E., da Costa, N.B., Desforges, J.E., Sánchez Schacht, J.R., Derry, A.M., Cooke, S.J., Barrett, R.D.H., Walsh, D.A., Ragoussis, J., Albert, M., Cristescu, M.E., & Gregory-Eaves, I. (2023). Prioritizing taxa for genetic reference database development to advance inland water conservation. Biological Conservation. https://doi.org/10.1016/j.biocon.2023.109963 | |
| Garcia-Elfring, A., Louchmanov, A.L., Sabin, C.E., Roffey, H.L., Samudra, S.P., Alcala, A.J., Lauderdale, J.D., Hendry, A.P., Menke, D.B., Barrett, R.D.H. (2023) Piebaldism and chromatophore development in reptiles is linked to the tfec gene. Current Biology. https://doi.org/10.1016/j.cub.2023.01.004 |
2022
| Glynn, V.M., Vollmer, S.V., Kline, D.I., Barrett, R.D.H. (2022) Environmental and geographic factors structure cauliflower coral’s algal symbioses across the Indo-Pacific. Journal of Biogeography. https://doi.org/10.1111/jbi.14560 | |
| Hu, J. and Barrett, R.D.H. (2022) Plastic and evolved DNA methylation during parallel adaptation of threespine stickleback (Gasterosteus aculeatus). Molecular Ecology. https://doi.org/10.1111/mec.16832 | |
| Martins, A.R.P., Martins, L.P., Ho, W., McMillan, W.O., Ready, J.S., Barrett, R.D.H. (2022) Scale-dependent environmental effects on phenotypic distributions in Heliconius butterflies. Ecology and Evolution. https://doi.org/10.1002/ece3.9286. | |
| Thurman, T.J., Palmer, T.M., Kolbe, J.J., Askary, A., Daskin, J.H., Gotanda, K.M., LaPierda, O., Kartzinel, T.R., Man in’t Veld, N.A., Revell, L., Wegener, H., Spiller, D.A., Losos, J.B., Pringle, R.M.P., Barrett, R.D.H. (2022) Predicting evolutionary change in response to novel ecological interactions: a field experiment with Anolis lizards. American Naturalist. https://doi.org/10.1086/723209 | |
| Wuitchik, S.J.S., Mogensen, S., Barry, T.N., Paccard, A., Jamniczky, H.A., Barrett, R.D.H.*, Rogers, S.M.* (2022) Evolution of thermal physiology alters the projected range of threespine stickleback under climate change. Molecular Ecology. https://doi.org/10.1111/mec.16396 | |
| Beausoleil, M., Camacho, C., Rabadán-González, J., Richard, R., Lalla, K., Carrion-Aviles, P., Hendry, A.P., Barrett, R.D.H. (2022) Where did the finch go? Insights from radio telemetry of the medium ground finch (Geospiza fortis). Ecology & Evolution. https://doi.org/10.1002/ece3.8768 |
2021
| Xu, C.C.Y., Ramsay, C., Cowan, M., Dehghani, M., Lasko, P., Barrett, R.D.H. (2021) Transgenes of genetically modified animals detected non-invasively via environmental DNA. PLOS ONE 16(8): e0249439. https://doi.org/10.1371/journal.pone.0249439 | |
| Hu, J., Wuitchik, S.J.S., Barry, T., Rogers, S.M., Jamniczky, H.A., Barrett, R.D.H. (2021) Heritability of DNA methylation in threespine stickleback (Gasterosteus aculeatus). Genetics 217: 1-15. | |
| Garcia‐Elfring, A., Paccard, A., Thurman, T.J., Wasserman, B.A., Palkovacs, E.P., Hendry, A.P. and Barrett, R.D.H. (2021), Using seasonal genomic changes to understand historical adaptation to new environments: Parallel selection on stickleback in highly‐variable estuaries. Molecular Ecology. https://doi.org/10.1111/mec.15879. | |
| Hébert, M.-P., Fugère V., Beisner, B.E., Costa N.B., Barrett R.D.H., Bell G., Shapiro B.J., Yargeau V., Gonzalez A., Fussmann G.F. (2021) Widespread agrochemicals differentially affect zooplankton biomass and community structure. Ecological Applications https://doi.org/10.1002/eap.2423 | |
| Costa N.B., Fugère V., Hébert M.-P., Costa N.B., Xu C.C., Barrett R.D.H., Beisner, B.E., Bell G., Fussmann G.F., Yargeau V., Fussmann, G.F., Gonzalez A., Shapiro B.J. (2021). Resistance, resilience, and functional redundancy of freshwater microbial communities facing multiple agricultural stressors in a mesocosm experiment. Molecular Ecology https://doi.org/10.1111/mec.16100. |
2020
| Fugère V., Hébert M.-P., Costa N.B., Xu C.C., Barrett R.D.H., Beisner, B.E., Bell G., Fussmann G.F., Shapiro B.J., Yargeau V., Gonzalez A. (2020) Community rescue in experimental freshwater ecosystems facing severe pesticide pollution. Nature Ecology and Evolution doi.org/10.1038/s41559-020-1134-5. | |
| Stange, M., Barrett, R.D.H., Hendry, A.P. (2020) Genetics and genomics for ecosystem services and nature’s contributions to people. Nature Reviews Genetics https://doi.org/10.1038/s41576-020-00288-7. | |
| Ecosystem size shapes antipredator trait evolution in estuarine threespine stickleback. Wasserman, B.A., Paccard, A., Apgar, T.M., Des Roches, S., Barrett, R.D.H., Hendry, A.P., Palkovacs, E.P. (2020) Oikos https://doi.org/10.1111/oik.07482. |
2019
2018
2017
2016
2015
2014
2013
2012
2011
2010
2009
2008
2006
2005
2004
Media
Popular Press
Please click on each entry to see stories about our research
Ho et al. Evolutionary Applications 2025
Xu et al. Current Biology 2025
Glynn et al. Current Biology 2025
Killarney Project
Xu et al. PRSB 2023
Sustainability in research: The Barrett lab on treading lightly in the field
Garcia-Elfring et al. Current Biology 2022
Bouncing Back in a Warming World
Xu et al. PLoS ONE 2021
Garcia‐Elfring et al. Molecular Ecology 2021
Fugere et al. Nature Ecology and Evolution 2020
Barrett et al. Science 2019
Pringle et al. Nature 2019
Interview on CBC’s The Current
Enclosure experiment in Nebraska
Linnen et al. Science 2013
Barrett et al. Proceedings of the Royal Society, Biological Sciences 2011
General stickleback press
Barrett et al. Science 2008
Barrett et al. Biology Letters 2006
Barrett et al. American Naturalist 2006
Research Posters



Conference Presentations
Outreach
The scientific community has a responsibility to communicate the importance of primary research to the public; unfortunately this is not always done effectively. As a result, we often fight an uphill battle against misconceptions and ignorance about basic scientific issues that we study. It is incumbent on scientists to make an effort to educate the public through outreach programs about the importance of our research for addressing basic and applied problems in science and society. To this end, we have made efforts to develop relationships with the communities where we conduct our fieldwork, and to contribute to science education through participation and organization of groups involved in public outreach. Below you can read about some of the fun initiatives Barrett lab members are engaged in.
I am the Redpath Museum’s graduate student Public Programs representative, where I strive to make the museum space a place where graduate student research is actively celebrated, and science accessible to visitors of all ages and backgrounds. Through this role, I am leading the curation of the Student research at the Museum exhibit, where each month we highlight a different student’s research and their trajectory in STEM, and begin to humanize science in the process. In particular, we are highlighting the voices of under-represented groups within STEM, as many people visiting museums feel that their stories and that of their communities are rarely showcased. This exhibit began as a physical installation in the second floor gallery, which has since transitioned online because of the COVID-19 pandemic. This work is done in close collaboration with the museum’s Public Programming administrative coordinator, Ms. Marie-Eve Leclerc. Current and past online exhibits can be accessed here.

I also enjoy giving talks to the general public about my research via the Museum. The Faculty of Science provides training on inquiry-driven outreach, where the goal is to co-construct knowledge with those in attendance, instead of assuming your audience is a “blank slate.” As such, my talks are very interactive, with constant questions from and to those tuned in, and visuals that the audience and I explore together. This process not only increases engagement, but also validates participants’ insights, subverting the science-public divide where scientists are positioned as the sole bearers of knowledge. One of my most memorable talks was my most recent one as part of the Québec-wide 24 heures de science festival and McGill’s 200th anniversary Family Science Day celebrations; you can read more about the outreach events that day in the following McGill Tribune article. Besides sharing with the general public the importance of coral reefs, what was most rewarding was hearing the impact my presentation had on the children in attendance. One of the children who came to my 24 heures de science talk drew me later that day during STEMM Diversity at McGill’s “draw a scientist” activity. Being able to share with younger generations that science takes many forms and that it is not a one-size-fits-all in terms of who can be a scientist, is why doing outreach is an integral part of my graduate studies at McGill.
-Victoria Marie Glynn


More than 10 scientists presented their work with art, theatre, animation and more at Ramène ta Science! Our lab participated in the My Thesis in 180 Seconds challenge to present our current work in speciation of Darwin’s finches and the importance of collaborative work in science. More specifically, we explained how new organisms can be created through the process of natural selection and how biodiversity is immensely important for ecosystem functioning and human populations. We were pleased to participate in a project making a direct connection between scientists and the public.
You can watch the video of the event here (in French):
https://www.facebook.com/ccstilarotonde.emse/videos/798827760282514/.
Link:
https://www.echosciences-loire.fr/articles/ma-science-en-180-secondes-par-marc-olivier.
I am also helping to develop the exhibit promoting diversity in STEM at the Redpath Museum. -Marc-Olivier Beausoleil
The importance of how science and scientists are perceived by the general public cannot be overstated. We often think of outreach as an extra thing on top of our research that – while is good to do, is ultimately somewhat optional. This paradigm needs to change. Science is a public endeavor supported by mostly public funds. We need justify our work and make it apparent that it is worth investing in for the good of everyone, especially in today’s political climate.
On April 20, 2017, I gave a public science talk to the Montreal Field Naturalists Club on how molecular genetics is revolutionizing the way we do conservation. I talked about how biologists use DNA from giant panda feces to estimate the number of wild pandas, which is partly the reason why they are no longer listed as an endangered species. I also talked about how environmental DNA of fish from feces, slime, etc. floating in lakes and rivers are being used to track invasive species of Asian Carp in the Mississippi River. Lastly, I talked about my research on using DNA from the bloodmeal of leeches to survey mammalian biodiversity in the Annamite forests of Vietnam and Laos.
https://montrealfieldnaturalists.wordpress.com/
I am also the founder of STEMM Diversity @ McGill, a student-drive initiative at the Redpath Museum to promote diversity in Science, Technology, Engineering, Math and Medicine. The initiative exists primarily as an online exhibit and a colouring/activity book. The online exhibit centers around interviews of diverse students and faculty about their personal experiences and opinions relating to the roles of gender and ethnicity in STEMM. These interviews were conducted by students and staff at the Redpath Museum in collaboration with TVM: Student Television at McGill. The exhibit also features various articles about diversity issues in STEMM as well as other student groups at McGill working towards greater diversity in STEMM. The exhibit is available at www.stemmdiversity.com/ and is now being used on 2 touchscreens in the second floor gallery of the Redpath Museum.
The STEMM Diversity @ McGill Colouring and Activity Book has received great reviews wherever we have sent it. The colouring book’s reach has expanded throughout and past the McGill community and continues to grow. To date, 275 colouring books have been distributed to elementary schools in Quebec and Ontario. Over 400 were distributed to eager participants of the NSERC Gender Summit 11. The colouring books have also been distributed to after school programs such as McGill’s Student-Parent Lunch and Learn, Homework Zone, which is an after-school program for underprivileged schools, Community and Family day, which was organized as part of McGill’s Black History Month, the Beaconsfield library, which does ample community engagement, and the entire Norfolk County library system in southern Ontario. 100 copies have gone to McGill’s series of Mini-Science lectures, and there are plans to send copies to Women in Resource Development Corporation’s STEM for GIRLS series. While colouring books are free to these educational institutions thanks to funds we have raised, the colouring books are also for sale in the Redpath Museum gift shop. All proceeds from the colouring book at the Redpath Museum go towards the museum’s outreach activities that encourage young people to pursue STEMM fields and their passions.
I also volunteer with “Lets Talk Science” (http://outreach.letstalkscience.ca/mcgill-university.html) to do science outreach to the general public. For example, we did an activity with visitors to the Redpath Museum where we blew carbon dioxide into red cabbage extract to simulate ocean acidification. -Charles Xu
I am currently involved in planning the Asokan Project, an initiative facilitated by Art History graduate student and member of Poundmaker Cree Nation Alexandra Nordstrom and myself in partnership with Chief Poundmaker School, which seeks to provide students from grades 5-9 of Chief Poundmaker School with summer programming focusing on the intersection between arts, culture, and science from an Indigenous perspective. We aim to engage students in collaborative learning that uses land as pedagogy for both scientific and artistic learning. Land as pedagogy is an ancient Cree concept that is linked with Indigenous identity and worldviews. This project is extremely important because it places an Indigenous worldview at the forefront of a collaborative learning process, creating a space for Indigenous youth to explore Cree culture and Indigenous ways of learning—an opportunity that is rarely offered in colonial and western institutions. Moreover, using art practice as a means to explore and communicate scientific learning brings together two disciplines that are generally thought of and taught as diametrically opposed in western schooling—arts and sciences. By bringing together arts and science, this project creates room for students to have sustainable and productive dialogues about how we can communicate environmental issues through art practice from an Indigenous point of view. Our project is aptly named the Asokan Project as “asokan” means bridge in Plains Cree and our main goal is to bridge gaps between arts and science, and Indigenous and western knowledge systems. We feel that it is necessary to bridge these gaps in order to be more productive and effective in any discipline and also to create welcoming spaces for everyone to participate in science. -Janay Fox
I had the great opportunity to work as an intern in the NGO Fundacíon Avifauna Eugene Eisenmann. The institution has as its main project the Panama Rainforest Discovery Center, an interactive nature center responsible for promoting Panamanian bird conservation, developing sustainable tourism and educational programs regarding sustainability, and working with the Panamanian community to administrate protected areas. During my internship I was involved in different sectors of the NGO, from administrative to daily maintenance tasks. One of the most satisfying moments was to take part in a project to raise emergence funds for the center, which is facing monetary difficulties and problems with floods and damaged trails. The center is a wonderful place to have an authentic experience of the lowland tropical rainforest and get in contact with environmental and conservation values. It worth knowing and contributing to the growth of this project. For more information: http://www.pipelineroad.org -Ananda Regina Pereira Martins
Join the Lab!
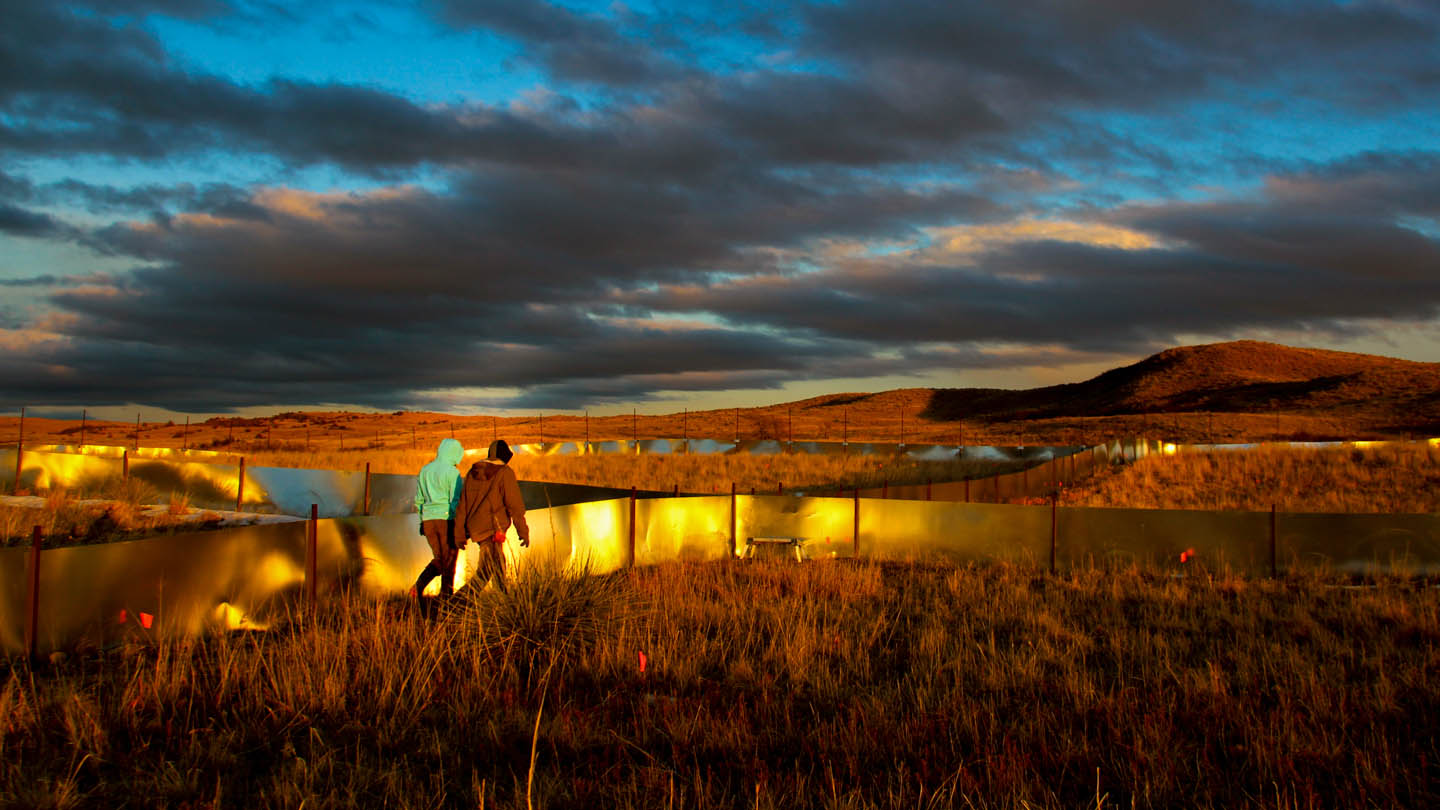

Graduate and Postdoctoral studies:
If you are interested in joining our lab group, please email me to introduce yourself. It would be most helpful if you could outline how your research interests overlap with mine and describe what you would be interested in working on during your time at McGill. Please also include your CV. To get a sense of our lab culture and resources, feel free to check out our continually evolving lab manual!
Benefits of working in our lab
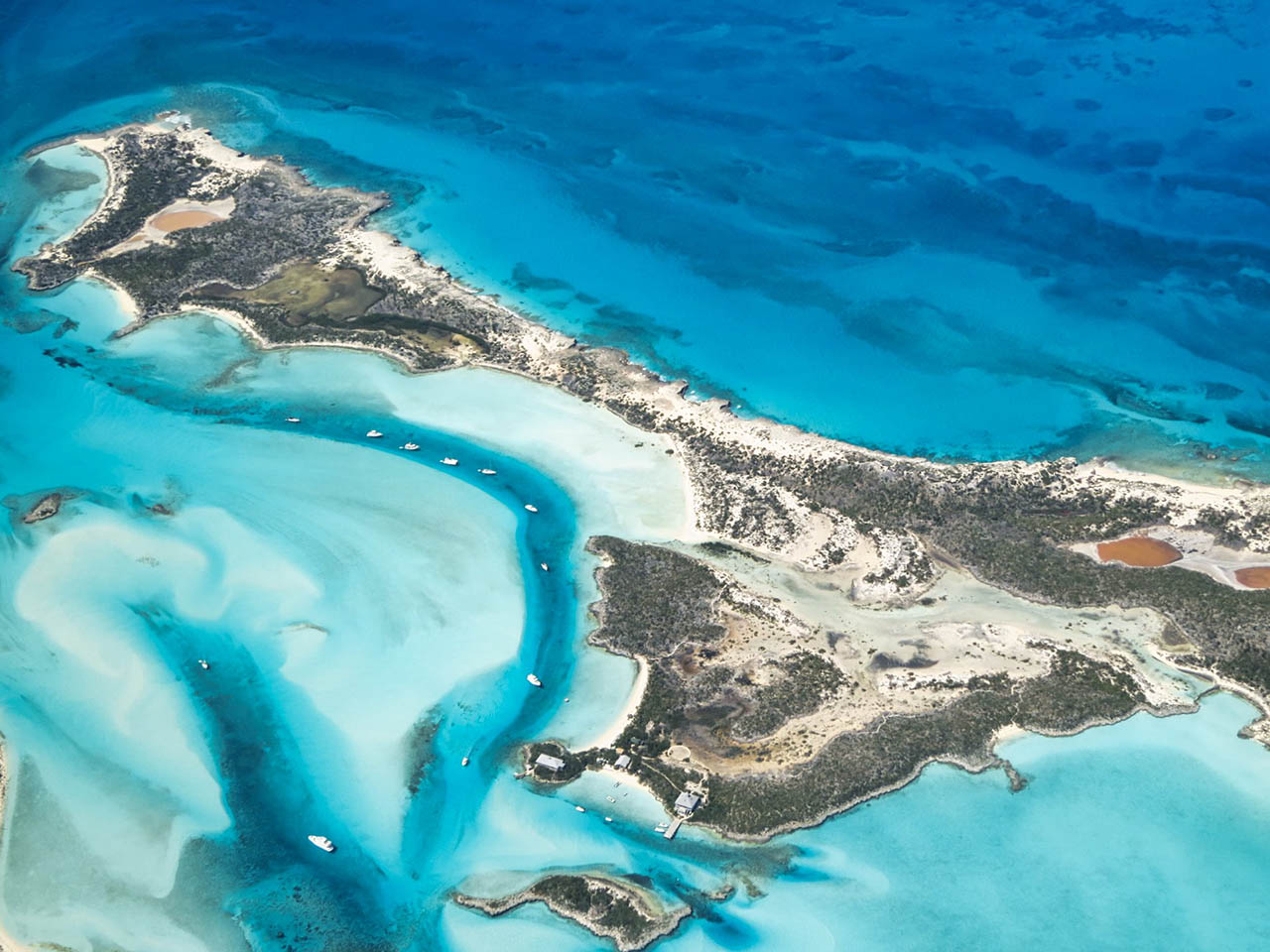
Amazing field work locations
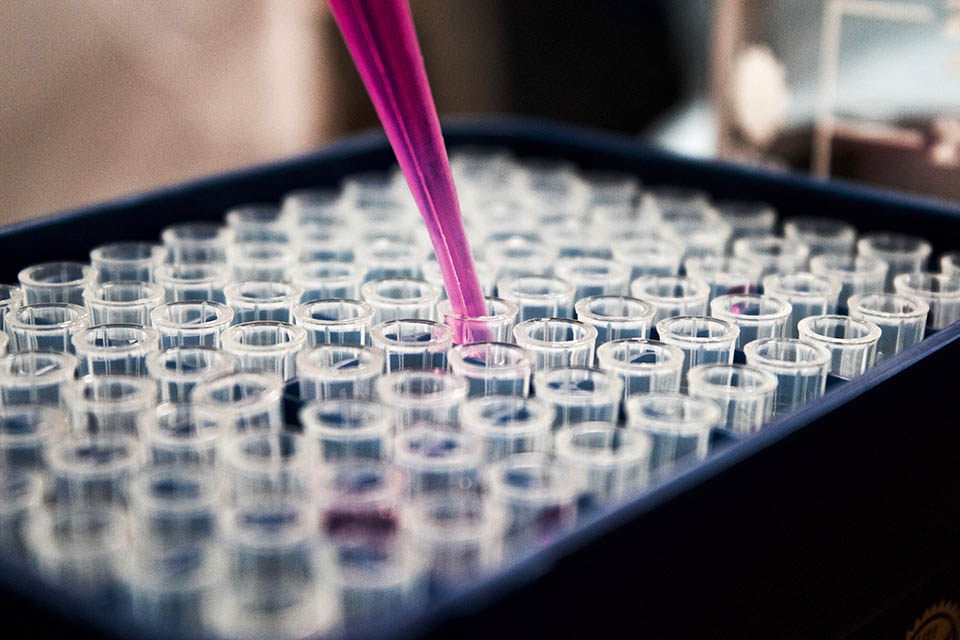


Cutting Edge Science



Sustainability
Barrett Lab is proud to receive a My Green Lab Platinum Certification.
Recognized by the United Nations Race to Zero campaign as a key measure of progress towards a zero-carbon future, My Green Lab Certification is considered the gold standard for laboratory sustainability best practices around the world.
My Green Lab Certification is a proven, scalable program that helps organizations achieve their sustainability goals. It offers tried-and-true methods rooted in science to dramatically reduce the environmental impact of laboratories without disrupting the critical work underway.
We are delighted to join a community of hundreds of labs that have been My Green Lab certified!.
More information about this program can be found at https://www.mygreenlab.org/green-lab-certification.html

Awards
- Winner of the 2023 International Freezer Challenge at McGill

Living in Montreal,
which is an awesome city!
Collaboration with a truly excellent and interactive group of people at the Biology Department
Benefits of working in our lab
Amazing field work locations
Cutting Edge Science
Living in Montreal,
which is an awesome city!
Collaboration with a truly excellent and interactive group of people at both the Redpath Museum and Biology Department
Student Resources
Funding
Here is a handy guide to help determine your eligibility for various forms of financial support.
And here is a link to a fees calculator for graduate tuition and fees for various types of degrees at McGill.
Note that students from Francophone countries may be eligible for international fee exemptions at McGill.
Here are a bunch of other funding sources I’ve come across (some for Canadians and others for international students/fellows):
Undergraduates
Are you an undergraduate student who wants to get a taste of scientific research? Please fill out this form. Below are a bunch of links with resources that could help you get paid while doing so.
Feel free to contact Wing or Lucas if you have any questions!
There are also many options for McGill undergraduate students to earn academic credits while doing research. Depending on your schedule, there are 1-, 3-, 6-, and 9-credit research course options. These can be undertaken by students in the Faculty of Science, Honours, Liberal or Joint programs, as well as students from the Faculty of Arts doing a Minor in Science. There is also the honours project option for committed students. For more information, please see: http://biology.mcgill.ca/undergrad/res_opps.html
And if you’re about to contact me to inquire about grad school, check this out!
International Students
A peek into our lab













































Marc-Olivier Beausoleil
Graduate Student


Janay Fox
Graduate Student
Janay completed a B.Sc. Honours in Genetics and a Certificate in Biological Research at the University of Alberta. As an undergraduate she used a candidate gene approach to investigate the genetic basis of boldness in a free ranging population of grey seals (Halichoerus grypus) under the supervision of Dr. Coltman. Her honours thesis project sought to predict the susceptibility of bighorn sheep (Ovis canadensis) to chronic wasting disease through sequencing of the prion protein gene, PRNP, under the supervision of Dr. Coltman and Dr. McKenzie. Janay is also passionate about bridging the gap between arts and science, and co-founded the asokan project with her colleague, Alexandra Nordstrom (MA Art History Concordia), which aims to create culturally relevant arts and science programming for Indigenous youth in Saskatchewan. For her PhD research she will be using the Trinidadian guppy system (Poecilia reticulate) to study how predation stress influences behavioural plasticity through epigenetics. Janay is part of the NEO-STRI program.


Alan Garcia-Elfring
Graduate Student
Alan obtained a BSc from Vancouver Island University. He then joined the Millien lab at McGill University, where he used genomic tools to investigate the evolutionary history of the white-footed mouse (Peromyscus leucopus) in southern Quebec. For his PhD, Alan is interested in using population genomics to investigate the genetic basis of phenotypic traits. In particular, he will focus on uncovering the genetic basis of recessive colour morphs in ball pythons (Python regius) found in herpetoculture.


Victoria Glynn
Graduate Student
Victoria earned her B.Sc. in Environmental Sciences from the University of California, Berkeley, minoring in Science and Math Education (CalTeach). As an undergraduate, she worked with Dr. Jamie Cate (UC Berkeley) using CRISPR-Cas9 to: genetically engineer yeast for specialized fatty-acid production, and reduce wheat and rice’s susceptibility to bacterial and fungal pathogens. Her honors thesis under Dr. Claire Kremen (UC Berkeley) studied how passerine and near-passerine birds in California’s Central Coast are impacted by agricultural land use change; this was achieved by linking haematological metrics and aerial imagery via mixed-effects modeling. Victoria is also passionate about STEM outreach. With the CalTeach program she designed culturally-relevant science curriculum and taught over 200 hours in K-12 classrooms in the East Bay. For her Ph.D. she is interested in exploring coral’s molecular responses to climate change. Victoria is part of the NEO-STRI Program and is being co-advised by Dr. David Kline (STRI).


Mary Kathleen (MK) Hickox
Graduate Student
MK obtained her BScH in Biological Sciences at Queen’s University and is a 2015 Schulich Leader Scholar. During her undergraduate studies, she studied evolution at both the behavioural (Dr. Adam Chippindale, Queen’s University) and genetic levels (Dr. Vicki Friesen, Queen’s University) to investigate speciation events. Her honours thesis, under the supervision of Dr. Steve Lougheed, investigated the population genomics of chorus frogs (Pseudacris maculata and P. triseriata) to help inform conservation practices. MK is an avid activist, and, in partnership with a non-for-profit NGO, has advocated to bring activism into academic spaces across the country. Her master’s thesis examines the micro-spatial distribution of Darwin’s finches on Santa Cruz Island, Galapagos. She is a proud member of the NEO program, and is studying under the co-supervision of Dr. Andrew Hendry.


Ananda Martins
Graduate Student
Ananda completed her BA in Biological Sciences at Federal University of Maranhão – Brazil (UFMA). As an undergraduate she worked on the systematics and phylogenetics of Arcas Swainson (Lycaenidae) and evolutionary aspects of male secondary sexual organs, under the supervision of Dr. Robert Robbins (National Museum of Natural History – Smithsonian Institution) and Dr. Gisele Garcia Azevedo (UFMA). She completed her M.Sc. at the Museum of Zoology – University of São Paulo/Brazil (MZUSP), working on the systematics and phylogenetics the Atlides Section (Lepidoptera: Lycaenidae, Theclinae, Eumaeini), under the orientation of Dr. Marcelo Duarte (MZUSP) and Dr. Robert Robbins (Smithsonian Institution). For her Ph.D. research, Ananda is interested in patterns of biodiversity and mechanisms that drive these patterns. She will focus especially on the process of hybridization, working with Heliconius butterflies of the Brazilian Amazon. Ananda is a member of the NEO program and is co-advised by Dr. Owen McMillan (STRI).


Mathilde Salamon
Graduate Student
Mathilde earned her BSc in Biological Sciences and a Magistère in Biology from Paris-Sud University (France). As an undergraduate, she worked on the mechanisms of genomic plasticity in the archea Thermococcus kodakarensis (supervised by Dr. Jacques Oberto, UPsud) and on the evolution of reproduction strategies in the ant Cataglyphis cursor (supervised by Dr. Thibaud Monnin, Pierre and Marie Curie University). She completed her MSc in Biodiversity, Ecology and Evolution (BEE) at Paris-Saclay University (France) after a one-year exchange at the University of Calgary. During her MSc, she worked on invasive species at the Roscoff Marine Station (France), studying trophic characteristics of native and invasive sympatric ascidian species of the genus Ciona (co-supervised by Dr. Christophe Lejeusne and Dr. Thierry Comtet), as well as connectivity patterns and population dynamics in the invasive kelp Undaria pinnatifida (supervised by Frédérique Viard). For her Ph.D, she is using genomic tools and resurrection ecology experiments to investigate the adaptation of a northern calanoid copepod (Leptodiaptomus minutus) to climate and historical acidification. Mathilde is based at the Université du Québec à Montréal and is co-supervised by Alison Derry. She is a member of the EcoLac training program.


Charles Xu
Graduate Student
Charles received his B.Sc. in environmental sciences from the University of Notre Dame. During his undergraduate, he worked on aquatic environmental DNA, speciation genomics of apple maggot flies, and spider web DNA. He then participated in the MEME Erasmus Mundus Masters Programme in Evolutionary Biology where he earned M.Sc. degrees from the University of Groningen, University of Montpellier, and Uppsala University. In his masters, he worked on the genetics of starvation tolerance in European seabass, population estimation of giant pandas using genetic mark recapture, and taxonomic assignment of metabarcoding data. Charles is broadly interested in the application of genetic tools towards answering questions and solving problems in ecology, evolution, and conservation. He also advocates for greater diversity and science outreach whenever possible. Charles was awarded a Vanier Canada Graduate Scholarship to support his Ph.D. on the discovery, study, and protection of biodiversity via metagenomics.


Åsa Lind
Research Associate
Åsa received her B.Sc. with Honours from the University of Glasgow in 2017. As an undergraduate, she conducted a project on population genetics in Atlantic salmon. After graduating, she worked as a technician at the Department of Integrative Biology at the University of Texas at Austin. During her time at UT, she worked on a meta-analysis on asymmetric premating isolation for Prof. Daniel Bolnick, conducted phonotaxis experiments on Túngara frogs for Prof. Michael J. Ryan and did ancient DNA work for Prof. Deborah Bolnick.


Wing-Zheng Ho
Research Associate
Wing majored in B.Sc Biology with a Specialization in Ecology at Concordia University. He is interested in developing new tools and frameworks to assist his peers in their ecological research and education. Currently volunteering at the Barrett lab, Wing is assisting with an experimental evolution project involving transplants of threespine stickleback into several Alaskan lakes. His current personal project consists of porting the Java based ecological modeling software Populus into an easily accessible web app for students without access to a compatible computer due to COVID-19 restrictions on computer lab access. App demo: wingzhg.shinyapps.io/populur. Wing’s website.


Tim Thurman
Graduate Student


Sara Wuitchik
Graduate Student
Sara conducted her Ph.D. in the Barrett and Rogers (University of Calgary) labs, investigating the ecological consequences of genetically-based thermal traits in fishes. She is currently a postdoctoral fellow with Michael Sorenson (University of Boston) and Tim Sackton (Harvard University), where she is working on the comparative genomics of avian brood parasitism.


Juntao Hu
Graduate Student
Juntao completed both his B.Sc. and M.Sc. in Agronomy at China Agricultural University. During his undergraduate and M.Sc., he worked on developing DNA barcoding techniques to identify invasive insect species, and explored molecular mechanisms, particularly heat shock protein expression, underlying the distinct range expansion patterns of two invasive fruit fly species. Juntao conducted his Ph.D. thesis in the Barrett lab, studying epigenetic responses to environmental change in wide range of animal taxa. Currently, he is a Postdoctoral Research Fellow at Harvard University, focusing on developing models to explore the relationship between DNA methylation, chronological age, and biological age in animals.


Arash Askary
Graduate Student
Ari investigated epigenetic responses to stress in Anolis sagrei during his Ph.D. He is currently doing market research for Hotspex.


Sara Kurland
Graduate Student
Sara was a visiting M.Sc. student in the lab. Her project focused on migration-selection balance in populations of threespine stickleback inhabiting bar-built estuaries in California. She is currently conducting her Ph.D. at the University of Stockholm.


Juliette Lemoine
Undergraduate Student (BIOL 479)
Juliette is completing her B.Sc in Biology at McGill University. For her Honours research project in BIOL 479, she worked with Charles Xu to investigate how the microbiome inhabiting skin piercings and how it changes over time after a disturbance event. She is using earlobe piercings as her study model to compare the microbiome of pierced and non-pierced skin from participants over time.


Savannah Bissegger-O’Connor
Undergraduate Student (BIOL 466)
Savannah is majoring in Biology at McGill University. She works in the lab as a research assistant performing sea anemone care and propagation. She is currently assisting Victoria Glynn in preparing a publication on the biogeography of Pocillopora corals and their dinoflagellates across the Indo-Pacific. For her Independent Research Project (BIOL 466), she studied the symbiotic relationship between the sea anemone Exaiptasia pallida and dinoflagellates. Her work involved determining the critical salinity point for our sea anemones and assessing their regenerative ability when exposed to induced asexual reproduction.


Frida-Cecilia Acosta-Parenteau
Undergraduate Student (BIOL 466)
Frida-Cecilia is majoring in Quantitative Biology at McGill University. For her Independent Research Project (BIOL 466), she worked with PhD student Victoria Glynn, and the undergraduate researcher Savannah Bissegger, studying the effects of water salinity and asexual reproduction in the emerging model system for corals, the sea anemone Exaiptasia pallida.


Avery Albert
Undergraduate Student
As an undergraduate, Avery worked closely with Charles in summer 2019 in order to help plan the “Microbiome of Human Piercings” project. Participants are currently being collected from a Tattoo Lounge in Montreal and we hope to begin bacterial sequencing of samples soon! Avery is currently completing a double major in the departments of Molecular Genetics & Microbiology and Ecology & Evolutionary Biology at the University of Toronto, as well as working in the lab of Dr. Aneil Agrawal in the EEB department.


Emily Brown
Undergraduate Student
Emily is a U2 Biology student at McGill. She volunteered with Ananda in 2019 and helped with standardized photography and digitization of the wings of Heliconius butterflies, and with finding patterns in wing colouration using R. Since her time in the Barrett lab she has worked on independent research in the von Sperber lab studying changes in phosphorus concentration in plants with varying altitude, bedrock type, and time in the growing season. Emily is currently studying in New Zealand for a semester, and hopes to continue to study human impacts on biodiversity in the future.


Nicole Stinson
Undergraduate Student
Nicole worked as a volunteer in Dr. Barrett’s lab during her undergrad, assisting Charles Xu with extracting environmental DNA from samples collected at the Large Experimental Array of Ponds (LEAP) to study community and evolutionary rescue. While working in the lab, she was able to practice DNA extractions, get comfortable with doing standard lab protocols such as PCR and gel electrophoresis. Nicole currently works for a non-profit, teaching and promoting science and robotics in schools in Kahnawake, and is planning to return to academia to complete a masters in evolutionary biology in the next few years.


Rachel Takasaki
Undergraduate Student
Rachel is a U2 biology student who worked on the DNA extractions from the 2017 Large Experimental Array of Ponds experiment, which used environmental DNA samples to study evolutionary and community rescue. Since leaving the Barrett lab, she has been working at the McGill Farmers’ Market and studying in Panama for a field study semester. She aspires to study the intersection of how ecosystems and human societies are adapting to climate change in the future.


Michael Maddalena
Undergraduate Student
In 2018, Michael was an undergraduate volunteer in the lab. He performed eDNA extractions from water samples collected at the Large Experimental Array of Ponds (LEAP) located at the Gault Nature Reserve to study the adaptability of the aquatic communities to acidity. Since then, Michael has also worked as an intern at INRS Armand-Frappier for Dr. Éric Déziel, studying the effects of multiple species and strains from the Burkholderia cepacia complex (BCC) on the growth of plants and the virulence of variant strains having lost a chromosome in a Galleria model. He is currently working towards his undergraduate degree in Microbiology and Immunology here at McGill.


Lauren Bennett
Undergraduate Student
Lauren worked in the Barrett lab in the spring and summer of 2018 assessing the effect of charged membrane filters in the collection of environmental DNA (eDNA) from natural water samples. In addition, Lauren helped extract eDNA from samples collected at the Large Experimental Array of Ponds (LEAP) facility in the summer of 2017 testing the effect of acidity on lake communities. She also participated in fieldwork for the LEAP 2018 experiment. Currently, Lauren teaches high school science to students with learning difficulties and continuously strive to make science exciting and accessible for all.


Scarlett (Yiyi) Xiao
Undergraduate Student
In summer 2018, Scarlett worked on measuring the surface area of leaves by using the Lamina program under the supervision of Tim Thurman and Charles Xu. These leaves were collected from experimental islands in the Bahamas and served as an indicator of herbivory. Scarlett is currently finishing her degrees in computer science and biology.


Kiran Yendamuri
Undergraduate Student
In the summer of 2017, Kiran assisted Charles Xu to develop an updated protocol for extracting DNA of spiders and their prey from spider webs. Kiran also worked with Charles as the director of videography for the STEMM Diversity @ McGill student initiative at the Redpath Museum. He has recently been accepted to the University of Manitoba to pursue an MSc in Arctic Oceanography and will start in the summer of 2020.


Samantha LaPenna
Undergraduate Student
Samantha worked in the Barrett Lab while completing her DEC in Health Science at Vanier College. She entered the BioGenius Competition in 2017 under the mentorship of Charles Xu. Her project was titled, “DNA extractions on spider webs vs DNA extractions from spider tissues; the next steps in conservation”. She is now in her final year at McGill University in Honours Life Sciences with a specialization in Animal Biology. She is currently conducting research in the Bordignon Lab on the Macdonald Campus. Her current project tests the efficiency of microinjection and electroporation in the delivery of CRISPR Cas9 and gene editing in porcine embryos. After graduation, she wishes to study veterinary medicine.


Natalie Warren
Undergraduate Student
Natalie is in her last semester at McGill with a B.Sc. in Environmental Biology. As an undergrad, she completed a research project in Dr. Jaqueline Bede’s Plant-Insect Interaction Lab, exploring how pine essential oil could act as a synergist, preventing the glutathione S-transferase detoxification mechanism in the cabbage looper caterpillar, Trichoplusia ni.


Jaren Mari Abergas
Undergraduate Student
Jaren is an undergraduate student at McGill University pursuing a B.Sc. in Microbiology & Immunology. He’s a recipient of the Light S.U.R.A. and will be researching in the Barrett Lab during the Spring and Summer of 2021. He is currently assisting the research of Graduate Student Alan Garcia on the genetic basis of color morph pigmentation of ball pythons, specifically on DNA extractions, quality control, quantification, and normalization.


Olivia Fraser Barsby
Undergraduate Student


Laura Lardinois
Graduate Student
Laura is fascinated by the trillions of microbes; including bacteria, archaea, viruses, and fungi, that grow on and inside living organisms, forming their microbiome. Her PhD research revolves around marine microbiomes and how they – in tandem with their animal hosts – can survive and adapt to both natural and anthropogenic environmental change. Her research leverages the seasonal upwelling in the Tropical Eastern Pacific and transisthmian sister fish species pairs in Panama’s coral reefs to investigate host-microbiome responses to changing environments across seasonal and evolutionary time scales. She is part of the NEO-STRI program and is co-supervised by Dr. Matthieu Leray at the Smithsonian Tropical Research Institute (STRI).
Laura completed her BA in Biology at Middlebury College (Vermont, USA). As an undergraduate, she was involved in various research projects including an investigation of cultural and geographic variation in personal care product (PCP) use worldwide with Dr. Kathryn Crawford (Middlebury College), and a study of the drivers behind seasonal shifts in aggression in a native electric teleost fish (Gymnotus omarorum) during an internship with Dr. Laura Quintana (IIBCE, Uruguay). She is passionate about science education and outreach, especially the use of art as a medium for science communication.


Lucas Eckert
Graduate Student
Lucas received his BSc from the Honours Integrated Science program at McMaster University, where he concentrated in Biology. As an undergraduate, he had the opportunity to work with Dr. Chad Harvey, Dr. Jim Quinn, and Dr. Sigal Balshine, researching various aspects of ecology and evolution. For his undergraduate thesis, Lucas worked with Dr. Sigal Balshine and Dr. Ben Bolker, using phylogenetic analysis to explore the evolution of reproductive accessory glands across fishes. For his Master’s thesis (co-supervised by Dr. Graham Bell), he will be using the stickleback system to investigate the predictability of evolution, through a large-scale transplant experiment in Alaska.


Eric Wootton
Graduate Student
Eric completed his honours BSc in Biochemistry and Molecular Biology at Trent University in 2022. Throughout his undergraduate degree, Eric was actively involved in various research projects focused on the population genetics of North American Ungulates as well as app development for science education. Eric received three NSERC Undergraduate Student Research Awards (USRAs) to work under Dr. Aaron Shafer (Trent University). For his undergraduate thesis, Eric investigated the genomic health of three demographically divergent ungulates: white-tailed deer, mountain goats, and caribou. He was also the co-president of the Undergraduate Chemistry Society at Trent University. In September 2022, Eric started his PhD studies with the Barrett lab at McGill University. His work focuses on the epigenetic modifications experienced by three-spined stickleback (Gasterosteus aculeatus) when the species is naturally subjected to changes in salinity. He is currently funded by an NSERC Canadian Graduate Scholarship (CGS).


Akhil Kholwadwala
Graduate Student
Akhil completed a B.S. in Ecology and Evolutionary Biology and a B.A. in French at the University of Rochester in 2022. As an undergraduate, Akhil conducted research with Dr. Jennifer Brisson (University of Rochester) on the evolution of polymorphic and polyphenic traits–such as color and wing morph–in aphids. He also conducted research with the United States Fish and Wildlife Service, testing the ability of eDNA technologies to detect the presence of rare and endangered bumblebee species. For his Ph.D. Akhil is co-supervised by Dr. Rowan Barrett and Dr. Jesse Shapiro. Using the LEAP mesocosm as a model, Akhil is interested in researching the adaptation of aquatic communities and individual species to a plurality of anthropogenic stressors and their synergistic effects. He is also interested in looking at the predictability of evolution within the context of this system.


Maxime Guglielmetti
Undergraduate Student
Maxime is an undergraduate student at McGill University majoring in Biology and minoring in Environment. In summer 2022 he worked with M.Sc. student Emma Derrick (co-supervised by Dr. Barrett) at the Large Experimental Array of Ponds (LEAP) lab as a Gault Research Award recipient. There, he looked at how biotic and abiotic factors affect the evolution of the bacteria in response to varying levels of glyphosate exposure. For his Honors project, he is working with Ph.D. student Victoria Glynn in the analysis of data she collected from coral reefs in Panama. They will try to resolve what microbiome composition and dynamics confers the most thermal resilience to coral hosts, and how environmental heterogeneity affects this resilience.


Linda Tang
Undergraduate Student
Linda is a fourth-year undergrad at McGill University majoring in Biology and minoring in Computer Science. For her honours thesis, Linda is working with Lucas Eckert to investigate rapid morphological changes in the three-spined stickleback in response to environmental change. She aspires to a career in education and to teach biology at the secondary school level.


Brook Brown
Undergraduate Student
Johanna Arnet
Undergraduate Student
Johanna is a fourth-year undergrad at McGill University majoring in Psychology with minors in Biology and Natural History. For an independent research project, Johanna is working with Lucas Eckert to investigate temporal morphometric changes in three-spine sticklebacks in experimental lakes in Alaska. She aims to continue in this field of biology and pursue graduate school.


Marcus Lin
Graduate Student
Marcus completed his BSc in Ecology, Evolution, and Behavior at the University of California Los Angeles. As an undergraduate, he worked with Dr. Tina Treude, Dr. David Jacobs, and Dr. Jeana Drake studying ocean geochemical cycles of nitrogen, coastal lagoon geomorphology, and stony coral biomineralization proteomics. Afterwards, Marcus worked with Dr. Sergey Nuzhdin at the University of Southern California, researching giant kelp (Macrocystis pyrifera) genetics and aquaculture. For his Master’s thesis (co-advised by Dr. Nagissa Mahmoudi), Marcus is interested in examining the genetics and microbial associations of local seaweed species under changing environments.


Alina Shimizu-Jozi
Undergraduate Student
Alina is an undergraduate student at McGill University entering her third year in the Joint
Honours Computer Science and Biology program. Alina recently attended a biology course held at McGill’s Bellairs Research institute in Holetown, Barbados. There, she pursued her interests in tropical biology by exploring size frequency distributions of coral species as a measure of coral reef health. Alina was also awarded a Summer Undergraduate Research in Engineering experience in collaboration with the Biofluids and Global Health Lab, lead by Dr. Caroline Wagner, pursuing research in infectious disease modeling and containment strategies. Continuing to explore her research interests, Alina will work with Ph. D student Victoria Glynn on her research in conducting heat stress experiments with sea anemones.


Andrew Kemp
Undergraduate Student
Andrew Kemp graduated from his Bachelor of Science in Honours Biology at McGill University in spring of 2024. He is broadly interested in evolutionary ecology, recently finishing his Honours Thesis with PhD student Lucas Eckert on morphological adaptations of stickleback along the benthic-limnetic axis in response to their environment, both in situ and in common garden. He has also done work on at-risk plants, peat soil, and biochar applications. His experiences have fostered his passion for researching climate change’s impacts on the environment and communities. The summer following his undergrad, he will be doing research using machine learning models trained on climatic datasets to help predict vulnerabilities and mitigate the effects of climate change in the Caribbean. He plans to pursue his interests through graduate studies.


Eva Savard
Undergraduate Student
Eva is an undergraduate student at McGill University, entering her final year in the Honours Biology program. She works as a volunteer in Dr. Barrett’s lab, assisting MK Hickox with extracting DNA from stickleback samples collected in California to study the processes of natural selection within and across populations. While working in the lab, she practiced various molecular techniques, such as DNA extractions, quantifications, or Speevac


Paige Smallman
Graduate Student


Tim Gemeinhardt
Graduate Student
Exploring a variety of systems and research questions, Tim pursued his BSc and MSc in Biochemistry at Goethe University, Germany. His projects led him to work on locomotion in C. elegans using optogenetic tools (Gottschalk lab, Goethe University), the piRNA pathway in the D. melanogaster germline (Kai lab, Osaka University), the transport mechanism of a bacterial membrane protein (Geertsma lab, Goethe University & Hummer lab, MPI Biophysics), and structural mass spectrometry to study membrane protein folding (Reading lab, King’s College London).
Tim’s exposure to diverse lines of research led to his broader interest in understanding the mechanisms and functions of epigenetics across length and time scales by bridging biochemistry and evolutionary biology. Tim is co-supervised by Dr Rowan Barrett and by Dr Nicole Francis at the Montreal Clinical Research Institute (IRCM), with whom he works on the molecular mechanisms of epigenetic regulators in the Polycomb Group. In collaboration with Dr Simon Reader, Tim is exploring epigenetic responses to predation stress in Trinidadian guppies. Besides his research projects, Tim is passionate to make science more sustainable, accessible and affordable.




Privacy Policy for Barrett Lab
At Barrett Lab, accessible from https://www.barrettlab.ca/, one of our main priorities is the privacy of our visitors. This Privacy Policy document contains types of information that is collected and recorded by Barrett Lab and how we use it.
If you have additional questions or require more information about our Privacy Policy, do not hesitate to contact us.
General Data Protection Regulation (GDPR)
We are a Data Controller of your information.
Barrett Lab legal basis for collecting and using the personal information described in this Privacy Policy depends on the Personal Information we collect and the specific context in which we collect the information:
- Barrett Lab needs to perform a contract with you
- You have given Barrett Lab permission to do so
- Processing your personal information is in Barrett Lab legitimate interests
- Barrett Lab needs to comply with the law
Barrett Lab will retain your personal information only for as long as is necessary for the purposes set out in this Privacy Policy. We will retain and use your information to the extent necessary to comply with our legal obligations, resolve disputes, and enforce our policies.
If you are a resident of the European Economic Area (EEA), you have certain data protection rights. If you wish to be informed what Personal Information we hold about you and if you want it to be removed from our systems, please contact us.
In certain circumstances, you have the following data protection rights:
- The right to access, update or to delete the information we have on you.
- The right of rectification.
- The right to object.
- The right of restriction.
- The right to data portability
- The right to withdraw consent
Log Files
Barrett Lab follows a standard procedure of using log files. These files log visitors when they visit websites. All hosting companies do this and a part of hosting services’ analytics. The information collected by log files include internet protocol (IP) addresses, browser type, Internet Service Provider (ISP), date and time stamp, referring/exit pages, and possibly the number of clicks. These are not linked to any information that is personally identifiable. The purpose of the information is for analyzing trends, administering the site, tracking users’ movement on the website, and gathering demographic information.
Cookies and Web Beacons
Like any other website, Barrett Lab uses ‘cookies’. These cookies are used to store information including visitors’ preferences, and the pages on the website that the visitor accessed or visited. The information is used to optimize the users’ experience by customizing our web page content based on visitors’ browser type and/or other information.
For more general information on cookies, please read “What Are Cookies”.
Privacy Policies
You may consult this list to find the Privacy Policy for each of the advertising partners of Barrett Lab.
Third-party ad servers or ad networks uses technologies like cookies, JavaScript, or Web Beacons that are used in their respective advertisements and links that appear on Barrett Lab, which are sent directly to users’ browser. They automatically receive your IP address when this occurs. These technologies are used to measure the effectiveness of their advertising campaigns and/or to personalize the advertising content that you see on websites that you visit.
Note that Barrett Lab has no access to or control over these cookies that are used by third-party advertisers.
Third Party Privacy Policies
Barrett Lab’s Privacy Policy does not apply to other advertisers or websites. Thus, we are advising you to consult the respective Privacy Policies of these third-party ad servers for more detailed information. It may include their practices and instructions about how to opt-out of certain options.
You can choose to disable cookies through your individual browser options. To know more detailed information about cookie management with specific web browsers, it can be found at the browsers’ respective websites.
Children’s Information
Another part of our priority is adding protection for children while using the internet. We encourage parents and guardians to observe, participate in, and/or monitor and guide their online activity.
Barrett Lab does not knowingly collect any Personal Identifiable Information from children under the age of 13. If you think that your child provided this kind of information on our website, we strongly encourage you to contact us immediately and we will do our best efforts to promptly remove such information from our records.
Online Privacy Policy Only
Our Privacy Policy applies only to our online activities and is valid for visitors to our website with regards to the information that they shared and/or collect in Barrett Lab. This policy is not applicable to any information collected offline or via channels other than this website.
Consent
By using our website, you hereby consent to our Privacy Policy and agree to its terms.